SUPG advection-diffusion of a Gaussian in an interval#
Donea, J. & Huerta, A. (2003). Finite Element Methods for Flow Problems. \(\S 5.6.1\)
\[\begin{split}
\mathbb{S}_u
\begin{cases}
\Omega = [0, 1] & \text{unit interval} \\
u_0(x) = \mu\exp\left(-(x-x_0)^2/\sigma^2\right) & \text{Gaussian initial condition} \\
u_{\text{D}}(x=0,1)=0 & \text{homogeneous Dirichlet boundary conditions} \\
\textbf{a}=a\,\textbf{e}_x & \text{constant velocity} \\
\mathsf{D}=D\mathsf{I} & \text{constant isotropic diffusivity} \\
R=0 & \text{zero reaction} \\
J=0 & \text{zero source} \\
u_{\text{e}}(x, t)=\frac{\mu}{\sqrt{1+4Dt/\sigma^2}}\exp\left(-\frac{(x-x_0-at)^2}{\sigma^2+4Dt}\right) & \text{exact solution} \\
Pe = \frac{a}{2DN_x} & \text{local Peclet number}
\end{cases}
\end{split}\]
import numpy as np
from lucifex.mesh import interval_mesh
from lucifex.fem import Constant
from lucifex.fdm import (
CN, advective_timestep, peclet_argument, FiniteDifference, FunctionSeries, ConstantSeries, finite_difference_order,
)
from lucifex.solver import ibvp , BoundaryConditions
from lucifex.sim import Simulation, run
from lucifex.plt import plot_line, save_figure
from lucifex.utils.npy_utils import as_index
from lucifex.utils.py_utils import nested_dict
from lucifex.pde.advection_diffusion import advection_diffusion
def exact_solution(
x,
t,
a,
d,
sigma,
mu,
x0,
):
factor = (mu / np.sqrt(1 + 4 * d * t / sigma**2))
func = np.exp(-(x - x0 - a * t)**2 / (sigma**2 + 4 * d * t))
return factor * func
def create_simulation(
tau: str | None,
Lx: float,
Nx: int,
dt: float,
a: float,
d: float,
D_adv: FiniteDifference,
D_diff: FiniteDifference,
mu: float,
x0: float,
sigma: float,
cache_matrix: bool = False,
) -> Simulation:
order = finite_difference_order(D_adv, D_diff)
mesh = interval_mesh(Lx, Nx)
t = ConstantSeries(mesh, name='t', ics=0.0)
dt = Constant(mesh, dt, name='dt')
a = Constant(mesh, (a, ), name='a')
d = Constant(mesh, d, name='d')
u = FunctionSeries((mesh, 'P', 1), name='u', order=order, store=1)
ics = lambda x: exact_solution(x[0], 0.0, a.value[0], d.value, sigma, mu, x0)
bcs = BoundaryConditions(
('dirichlet', lambda x: x[0], 0.0),
('dirichlet', lambda x: x[0] - Lx, 0.0),
)
u_solver = ibvp(advection_diffusion, ics, bcs, cache_matrix=cache_matrix)(
u, dt, a, d, D_adv, D_diff, tau=tau,
)
return Simulation(u_solver, t, dt)
Lx = 1.0
Nx = 100
h = Lx / Nx
a = 1.0
courant = 0.5
dt = advective_timestep(a, h, courant)
D_adv = CN
D_diff = CN
mu = 5/7
x0 = 2/15
sigma = 7 * np.sqrt(2) / 300
Pe_opts = (1.0, 5.0, 100.0)
tau_opts = (None, 'coth', 'shakib', 'codina')
simulations = nested_dict((str, float, Simulation))
for tau in tau_opts:
for Pe in Pe_opts:
d = peclet_argument(Pe, h=h, a=a)
simulations[tau][Pe] = create_simulation(tau, Lx, Nx, dt, a, d, D_adv, D_diff, mu, x0, sigma)
t_stop = 0.7
for tau in tau_opts:
for Pe in Pe_opts:
run(simulations[tau][Pe], t_stop=t_stop)
x = np.linspace(0, Lx, num=500)
t_target = 0.6
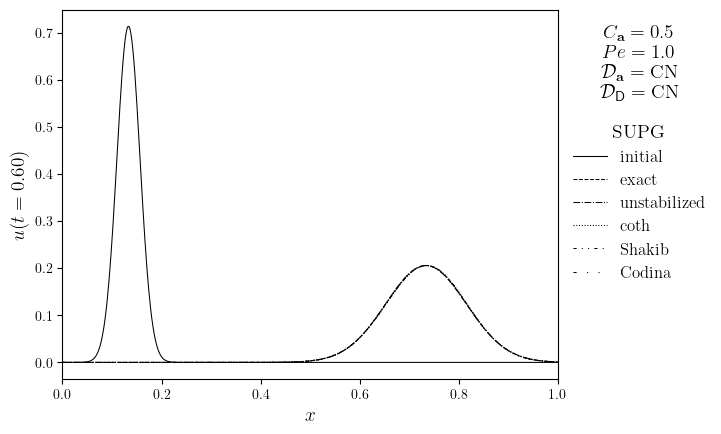
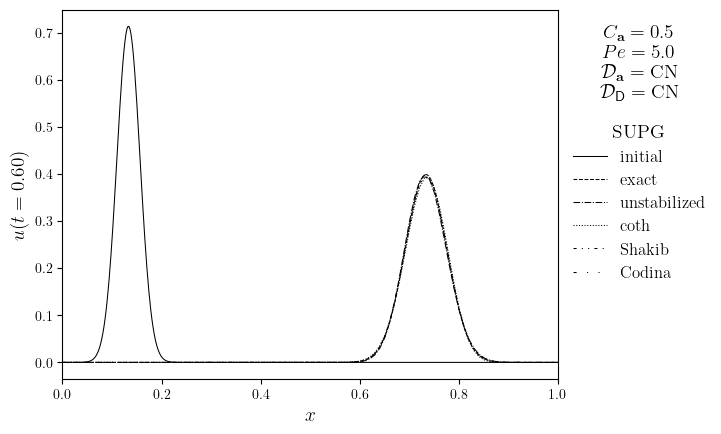
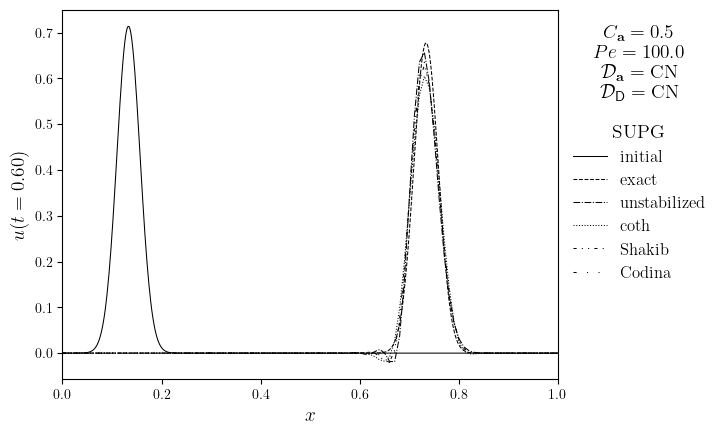
for Pe in Pe_opts:
legend_title = '\n'.join(
(
f'$C_{{\mathbf{{a}}}}={courant}$',
f'$Pe={Pe:.1f}$',
f'$\mathcal{{D}}_{{\mathbf{{a}},u}}=\mathrm{{{D_adv.name}}}$',
f'$\mathcal{{D}}_{{\mathsf{{D}},u}}=\mathrm{{{D_diff.name}}}$',
'\nSUPG',
)
)
d = peclet_argument(Pe, h=h, a=a)
ue = exact_solution(x, t_target, a, d, sigma, mu, x0)
u0 = exact_solution(x, 0.0, a, d, sigma, mu, x0)
lines = [(x, u0), (x, ue)]
legend_labels = ['initial', 'exact']
for tau in tau_opts:
u = simulations[tau][Pe]['u']
time_index = as_index(u.time_series, t_target)
t = u.time_series[time_index]
lines.append(u.series[time_index])
if tau is None:
tau = 'unstabilized'
if tau in ('shakib', 'codina'):
tau = tau.capitalize()
legend_labels.append(tau)
fig, ax = plot_line(
lines, legend_labels, legend_title,
x_lims=x,
x_label='$x$',
y_label=f'$u(t={t:.2f})$',
)
fig_name = f'Pe={Pe:.1f}_C_{{\mathbf{{a}}}}={courant}_Dadv={D_adv.name}_Ddiff={D_diff.name}'
save_figure(fig_name, thumbnail=(Pe is Pe_opts[-1]))(fig)
The Kernel crashed while executing code in the current cell or a previous cell.
Please review the code in the cell(s) to identify a possible cause of the failure.
Click <a href='https://aka.ms/vscodeJupyterKernelCrash'>here</a> for more info.
View Jupyter <a href='command:jupyter.viewOutput'>log</a> for further details.